- Subject:
- Applied Science
- Biology
- Life Science
- Material Type:
- Module
- Date Added:
- 07/10/2017


By the end of this section, you will be able to:Identify and describe the properties of lifeDescribe the levels of organization among living thingsRecognize and interpret a phylogenetic treeList examples of different sub disciplines in biology

By the end of this section, you will be able to:Identify and describe the properties of lifeDescribe the levels of organization among living thingsRecognize and interpret a phylogenetic treeList examples of different sub disciplines in biology

This resource is a video abstract of a research paper created by Research Square on behalf of its authors. It provides a synopsis that's easy to understand, and can be used to introduce the topics it covers to students, researchers, and the general public. The video's transcript is also provided in full, with a portion provided below for preview:
"Microbiomes are more than just prokaryotes and viruses; they also contain important eukaryotes, including fungi and protists. However, eukaryotes are difficult to study using ‘shotgun’ metagenomics, as their signal is often overwhelmed by the prokaryotes. Some methods use eukaryote-specific marker genes, but they can’t detect eukaryotes that aren’t in the reference marker gene set, and such methods are not compatible with web-based tools for downstream analysis. But CORRAL (Clustering Of Related Reference ALignments) is designed to close those gaps. CORRAL identifies eukaryotes in metagenomic data based on alignments to eukaryote-specific marker genes and Markov clustering. It can detect microbial eukaryotes that are not included in the marker gene reference set. The process is even automated and can be carried out at scale. A recent paper demonstrates CORRAL’s sensitivity and accuracy with simulated datasets, mock community standards, and human microbiome datasets..."
The rest of the transcript, along with a link to the research itself, is available on the resource itself.
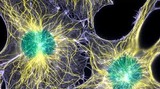
In this lesson, the students look at the components of cells and their functions. The lesson focuses on the difference between prokaryotic and eukaryotic cells. Each part of the cell performs a specific function that is vital for the cell's survival. Bacteria are single-celled organisms that are very important to engineers. Engineers can use bacteria to break down toxic materials in a process called bioremediation, and they can also kill or disable harmful bacteria through disinfection.

In this unit, students look at the components of cells and their functions and discover the controversy behind stem cell research. The first lesson focuses on the difference between prokaryotic and eukaryotic cells. In the second lesson, students learn about the basics of cellular respiration. They also learn about the application of cellular respiration to engineering and bioremediation. The third lesson continues students' education on cells in the human body and how (and why) engineers are involved in the research of stem cell behavior.

This resource is a video abstract of a research paper created by Research Square on behalf of its authors. It provides a synopsis that's easy to understand, and can be used to introduce the topics it covers to students, researchers, and the general public. The video's transcript is also provided in full, with a portion provided below for preview:
"Genomics research has greatly increased understanding of the human gut microbiome, but the existing reference databases remain insufficient, failing to map up to half of the sequences obtained in human gut studies. To solve this problem, researchers recently created HumGut, a comprehensive global reference database for the genomes of gut microbes in healthy humans. The researchers built the database by comparing nearly half a million publicly available prokaryote genomes with over 3,500 gut metagenomes from healthy humans worldwide, and retaining the prokaryote genomes that closely matched the sequences in healthy human guts. HumGut was approximately the same size as the recently released UHGG collection and half the size of a standard reference database. However, HumGut outperformed both other databases in classifying metagenomic reads from human gut samples, resulting in a lower percentage of unclassified reads..."
The rest of the transcript, along with a link to the research itself, is available on the resource itself.
Our second video from the cell biology lesson, part of our anatomy and physiology lecture series.
This video gives a brief summary of the differences between eukaryotes and prokaryotes.
All of our videos can be found at http://www.mrfordsclass.net
The concepts covered in this video include:
•Eukaryotes
•Prokaryotes
Students culture cells in order to find out which type of surfactant (in this case, soap) is best at removing bacteria. Groups culture cells from unwashed hands and add regular bar soap, regular liquid soap, anti-bacterial soap, dishwasher soap, and hand sanitizer to the cultures. The cultures are allowed to grow for two days and then the students assess which type of soap design did the best job of removing bacteria cells from unwashed hands. Students extend their knowledge of engineering and surfactants for different environmental applications.

This course covers introductory microbiology from a systems perspective, considering microbial diversity, population dynamics, and genomics. Emphasis is placed on the delicate balance between microbes and humans, and the changes that result in the emergence of infectious diseases and antimicrobial resistance. The case study approach covers such topics as vaccines, toxins, biodefense, and infections including Legionnaire’s disease, tuberculosis, Helicobacter pylori, and plague.

This resource is a video abstract of a research paper created by Research Square on behalf of its authors. It provides a synopsis that's easy to understand, and can be used to introduce the topics it covers to students, researchers, and the general public. The video's transcript is also provided in full, with a portion provided below for preview:
"The termite gut is the world’s smallest bioreactor and the most efficient system for breaking down biomass. To learn how this mini-digester might one day be scaled up to a technologically meaningful level, researchers examined the structure and function of the gut microbiomes from 11 termite genera which were grouped by diet into plant-fiber feeders and soil feeders. Both groups had similar bacterial flora. But subtle differences did emerge, with each termite species harbouring a unique set of genes encoding for breaking down plant biomass. Future metagenomics studies could help refine the specific functions of different bacterial genes within the termite gut, allowing for better insight into the termite–bacteria relationship and teasing out capabilities that could help bring these microscopic reactors to the macroscale..."
The rest of the transcript, along with a link to the research itself, is available on the resource itself.